Functional Simulation with Advanced Point Neurons (E-GLIF)¶
This Use Case can be used to simulate an olivocerebellar microcircuit with point neurons modeled as E-GLIF (Extended-Generalized Leaky Integrate and Fire). This advanced point neuron model has been designed and optimized to reproduce complex electroresponsive dynamics (including bursting, adaptation, resonance, etc) while keeping a limited computational load [Geminiani et al., 2018]. Simulations are run in NEST 2.18 and show how realistic single neuron dynamics and network topology contribute to network spiking patterns following sensory-like stimuli.
The Use Case can be found in Online Use Cases/Brain area circuit in silico experiments/Cerebellum/Functional Simulation with Advanced Point Neurons (E-GLIF).
Simulation description
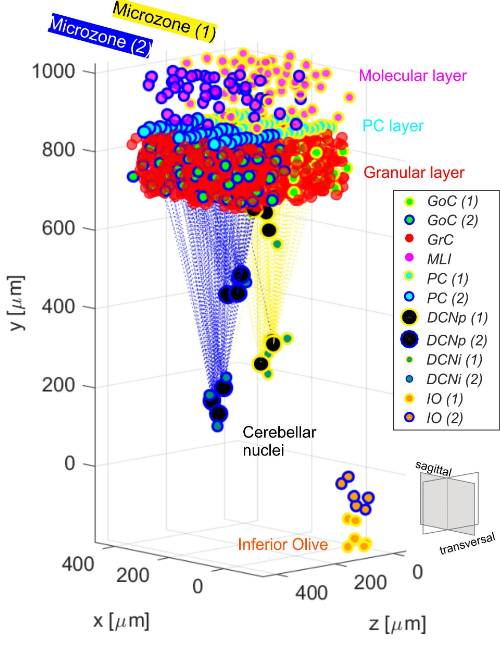
The olivocerebellar Spiking Neural Network is created in pyNEST. The topology is reconstructed based on the cerebellar scaffold rules in [Casali et al. 2019], adding the Olivary nuclei and a subdivision into microcomplexes to the original scaffold. Neurons are modelled as E-GLIF with alpha-shaped conductance-based synapses [Geminiani et al., 2018]. E-GLIF parameters are specific for each cell type, as optimized in [Geminiani et al., 2019]. Connection delays are obtained from literature values, while connection weights are tuned with a trial and error procedure to obtain physiological baseline population firing rates. The network is stimulated with sensory-like input signals to the Granular layer and the Inferior Olive, mimicking the input signals of Eye-Blink Classical Conditioning: a constant Poisson spike pattern at 40 Hz to the Granular layer, and a burst at 500 Hz to one nucleus of the Inferior Olive. Realistic single neuron dynamics provided by the E-GLIF and modular network topology allow to reproduce the complex spiking patterns of cerebellar signal encoding, including burst-pause (in Purkinje cells) and pause-burst (in Deep Cerebellar Nuclei cells) mechanisms.
Inputs
a single hdf5 file (saved in /storage) containing network topology data, i.e. cell positions and connections.
The user can modify the stimulation protocol by changing the parameters in the “Defining stimuli” section, namely: the duration of the simulation [ms]; the start [ms], end [ms] and frequency [Hz] of each stimulus; the radius [µm] of the stimulated area in the Granular layer.
Output
Files: .gdf files containing spike times of each network neuron over the simulation duration. The user can also choose to save the .dat files containing the voltage traces (but this would significantly increase the simulation time if whole population are selected)
Monitoring: raster plot of spikes for each neuron type, which the user can select, and PSTH
Additional information
The whole Use Case should take about 40 minutes to be completed, for a volume base of 400 x 400 µm.
The BSP foundation software used in the notebook is pyNEST
Single neuron and network parameters are the same reported in [Geminiani et al., 2019]
EXAMPLE
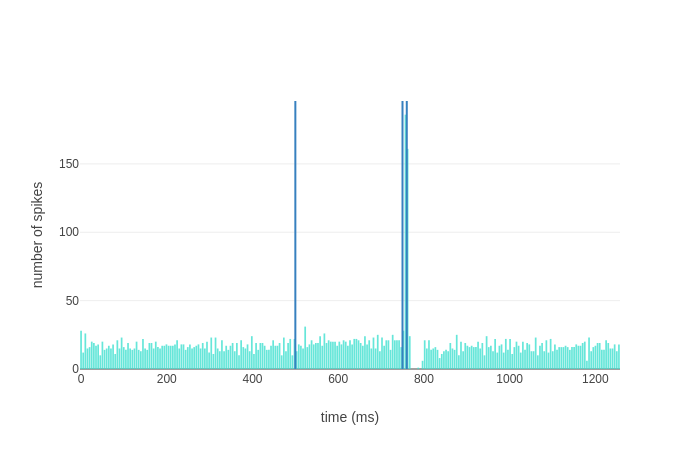
Providing the olivocerebellar scaffold a Poisson spike train (40 Hz frequency, 260 ms duration) to the Granular layer and a co-terminating burst (500 Hz frequency, 10 ms duration) to the Olivary nucleus connected to the first microcomplex, the following PSTH should be generated for Purkinje cells in the first microcomplex (vertical lines representing start of first and second stimulus, end of both stimuli):
with the first stimulus causing an increased population firing rate and the burst causing a complex spike (burst-pause).
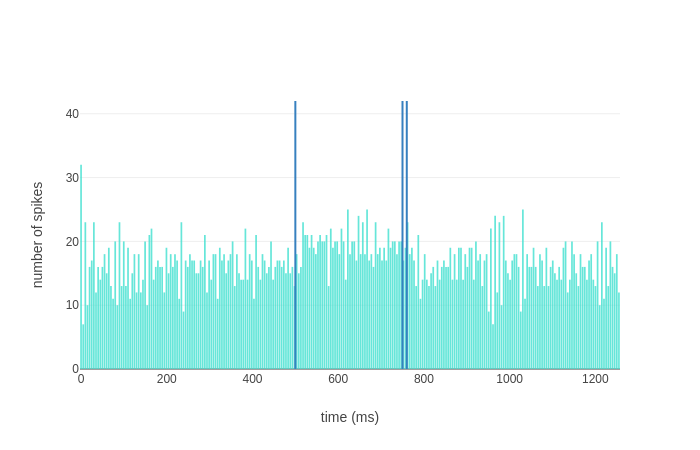
The bursting spiking pattern is not present in the Purkinje cells of the second microcomplex, which receive only the first stimulus thanks to the modular connectivity of the network:
with the complex spike not occurring.