Single cell in silico experiments under current clamp¶
The Single cell in silico experiments under current/voltage clamp Use Case provides Web UI to access Neuron as a Service in order to load the single cell models and run model simulations.
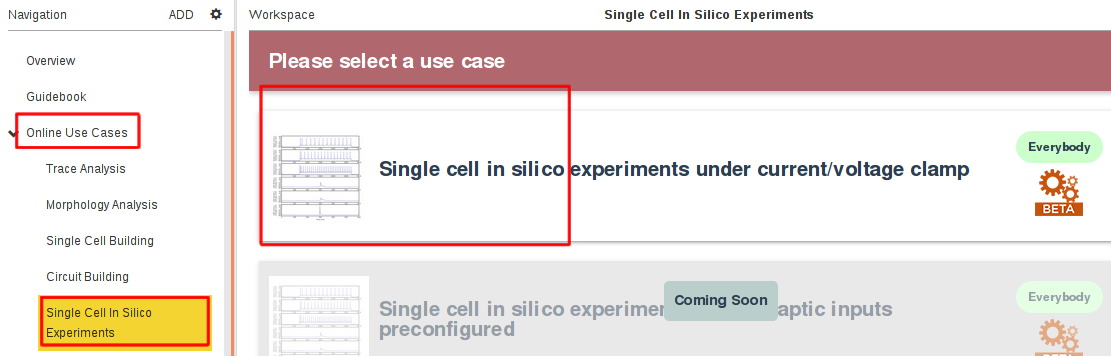
This Use Case can be run through the Online Use Cases/Single Cell In Silico Experiments panel in the Brain Simulation Platform (click here to access the tool):
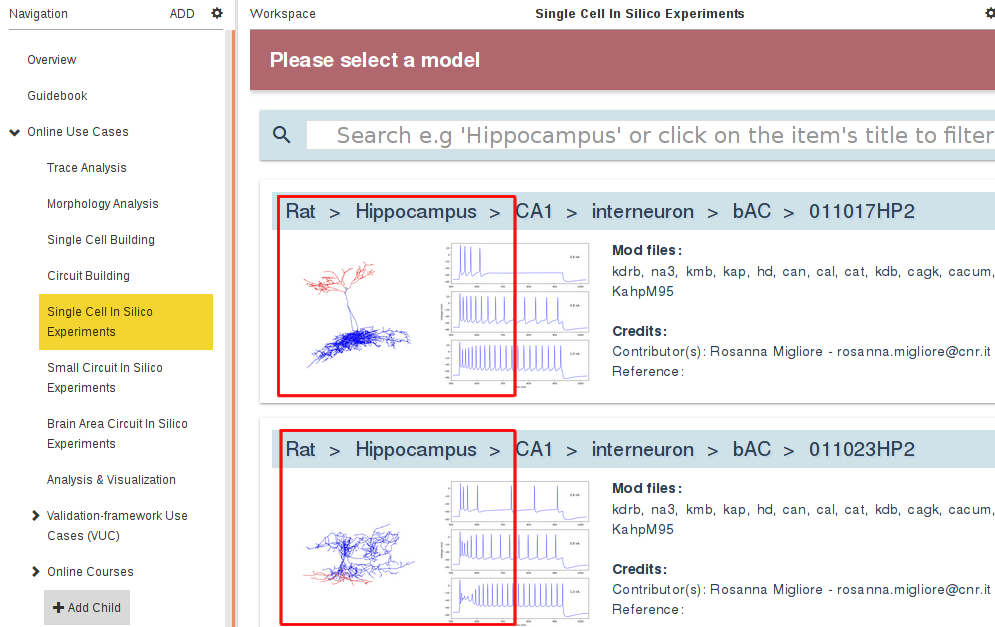
When you arrive at the Single cell in silico experiments under current/voltage clamp Use Case you can select the single cell model from the HBP data:
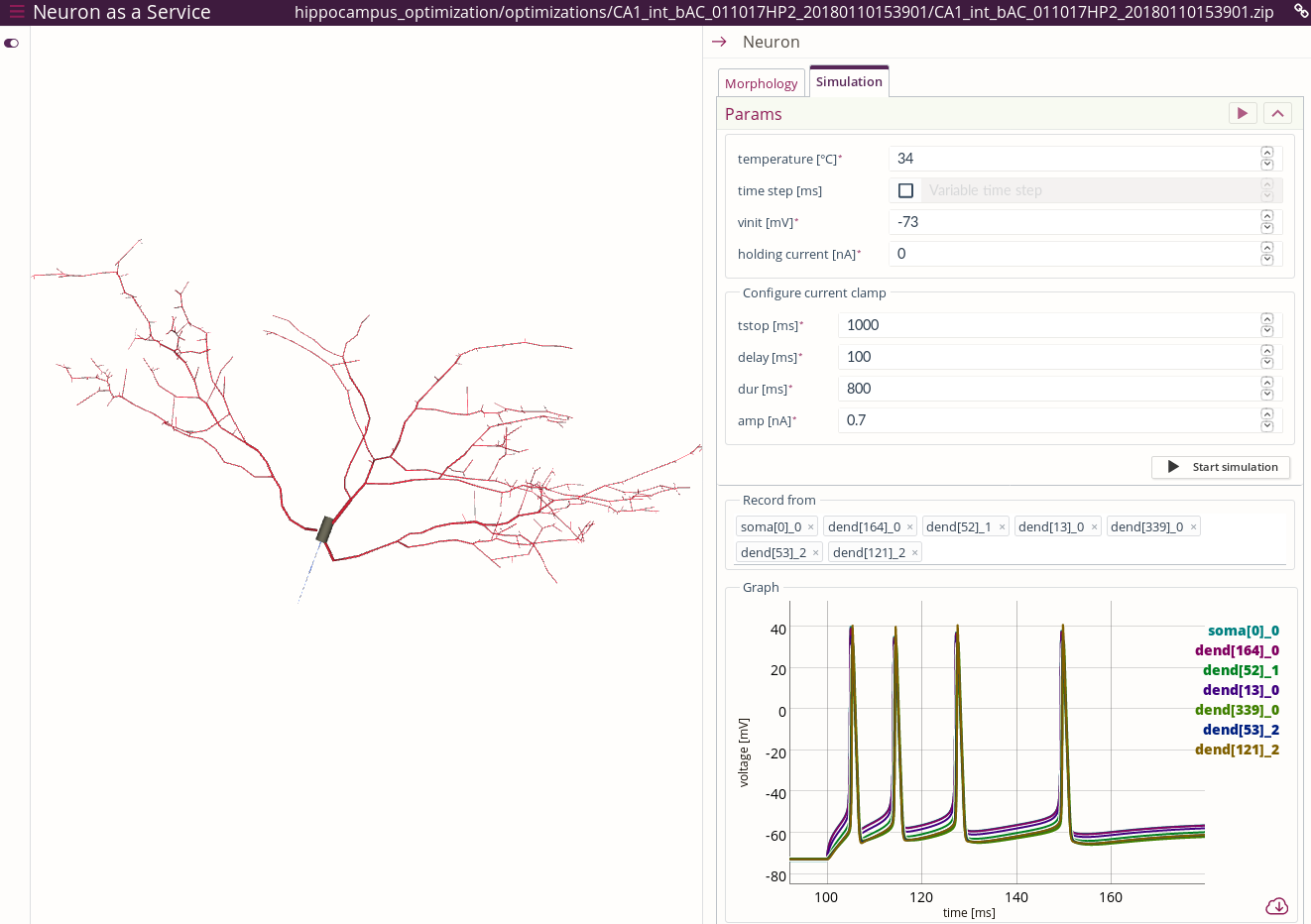
As a result, the single cell model will open in a new Browser window within the Neuron as a Service web application. The following screenshot of the app is annotated with red text to guide you through the interface:
Cell morphology can be visualized either in 3D view or 2D Dendrogram view. It is possible to zoom/rotate/pan using your mouse/scroll wheel/trackpad.
Current clamp can be attached to any section by clicking the segment in the cell 3D/2D Dendrogram view and then pressing the Place current injection button, or by selecting the section in the tree view on the right and pressing the same button.
To run the simulation, switch to the Simulation tab. In this view, you can click and select the segments to record the voltage from. They are added to the corresponding list. Up to 10 segments can be recorded from.
The graph showing the traces recorded from the cell segments can be zoomed in by clicking and dragging in order to select the area to zoom in. Double click on the graph will restore the original zoom level.
The recorded traces can be downloaded as csv file. The download link is available at the bottom right corner of the graph after the simulation has finished.
The following jupyter notebook code shows how it can be loaded with pandas for the further analysis:
import pandas as pd %matplotlib inline df = pd.read_csv('sim_CA1_int_bAC_011023HP2_20170510120324_2017-06-21_14-36-10_amp-soma_0-0.7nA.csv') df.plot.line(x='time', y='soma[0]_0')
In order to initialize simulation parameters with certain values it is possible to append the application URL with, for example, the following:
?delay=0&vinit=-86&dt=.025&=0&hypamp=0&tstop=500&delay=100&dur=800&celsius=37If the model folder contains file named
traces.datcontaining experimental traces in the following format:time,soma[0]_0_exp,soma[0]_1_exp,soma[0]_2_exp,soma[0]_3_exp,soma[0]_4_exp,soma[0]_5_exp 0.004,-87.179,-87.136,-86.605,-86.825,-86.453,-85.763 0.044,-87.249,-87.053,-86.521,-86.863,-86.425,-85.787 0.084,-87.254,-87.013,-86.503,-86.861,-86.445,-85.776
Then traces from this file will be loaded after the simulation completes, and in addition to the simulation traces. If trace names following the naming convention:
SECTION\_SEGMENTINDEX\_ANYNAMEthen the corresponding segment will be highlighted when user hovers the mouse over the trace.If the model folder contains file named
synapses_meta.jsonin the following format:{ "exc": ["AMPANMDA_EMS"], "inh": ["GABAAB_EMS"] }
Then those channel mechanisms will be visualized on the morphology with the corresponding coloring for excitatory and inhibitory.
If you have spines modeled as additional sections, it is assumed that they are constructed from a single segment. They will also be correctly positioned and visualized on the morphology.