Preparing a protein to run an atomistic Molecular Dynamics simulation¶
Overview¶
This use case aims to illustrate the process of setting up an atomistic Molecular Dynamics simulation system containing a protein, step by step. The particular example used is the Parvalbumin Calcium-binding protein (PDB code 1B8R).
Parvalbumin is a calcium binding protein that is found in subsets of sensory neurons, inhibitory interneurons and motoneurons. Alterations in the function of parvalbumin-expressing neurons have been implicated in various areas of clinical interest such as Alzheimer’s disease, autism, schizophrenia, age-related cognitive defects and some forms of cancer.
Background¶
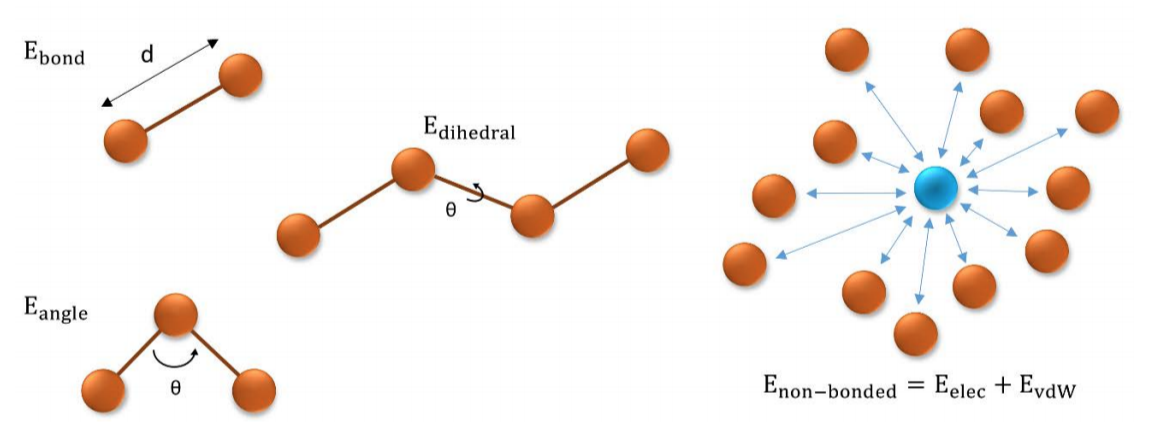
Molecular Dynamics (MD) simulation is the most popular theoretical technique to obtain macromolecular dynamic information. Classical mechanics is used to represent atoms as spheres of a given radius, hardness, charge and mass. The energy functional used by force-fields is usually composed of two terms: bonded and non-bonded components:
where
and
The combination of the force-fields with the laws of classical mechanics (Newton’s second law of motion), allows the calculation of the time evolution of the system. Trajectories of atoms and molecules are determined by numerically solving Newton’s equations of motion for a system of interacting particles, where forces between the particles and their potential energies are calculated using the force-field energy functionals.
How it works ?¶
This workflow makes extensive use of the BioExcel Building Blocks library (biobb). Each step of the process is performed by a building block (bb), which are wrappers of tools/scripts that computes a particular functionality (e.g. Solvating a system). If you are interested in expanding/modifying the current workflow, please visit the existing documentation for each of the packages here.
Most of the steps performed in this pipeline run GROMACS MD package tools, one of the most popular MD packages available.
Although the pipeline is presented step by step with associated information, it is extremely advisable to previously spend some time reading documentation about Molecular Dynamics simulations, to get familiar with the terms used, especially for newcomers to the field. This workflow is based on the official GROMACS MD setup tutorial: http://www.mdtutorials.com/gmx/lysozyme/index.html
Outcomes / Steps¶
This use case will explain:
How to fetch a PDB structure from the RCSB PDB database
How to fix a protein structure (add missing atoms)
How to create a protein system topology
How to create a solvent box surrounding the protein
How to fill the box with water molecules
How to energetically neutralize the system with the addition of counterions
How to energetically minimize the system
How to equilibrate the system in two steps (NVT, NPT)
How to run a free Molecular Dynamics (MD) simulation
How to post-process and visualize the resulting 3D trajectory
How to find and download the generated output files
Input¶
A PDB code of a protein structure
Outputs¶
Interactive and 3D vizualisation of the intermediate results on the protein structure
Interactive and 3D vizualisation of the resulting trajectory
Interactive visualization of 2D analyses plots (energies, temperature, RMSd)
Short (100ps) trajectory file generated from the final free MD simulation step
Collection of files needed to extend the MD simulation available to download
Targeted audience¶
All scientists working in biology related areas where protein study is relevant with a focus on structural biologists and biochemists. Especially directed to scientists interested in protein dynamics and flexibility.