Synaptic plasticity pathways¶
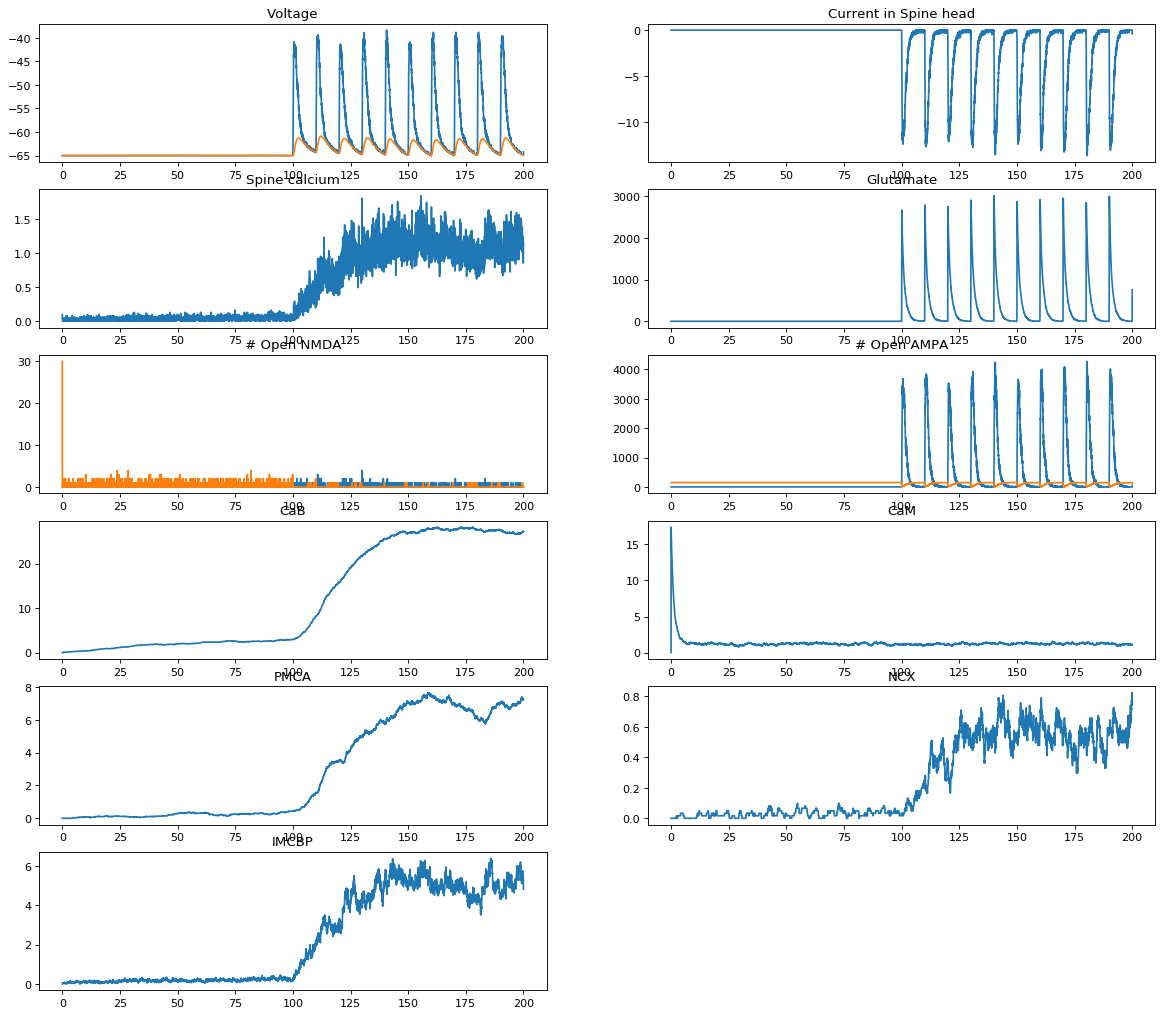
As discussed in the previous section, the motivation is to model the interaction between electrical activity in neurons and the synaptic proteome. This example (currently at https://collab.humanbrainproject.eu/#/collab/2184/nav/303192) shows how interactions between a number of ions, proteins and second messengers can be modeled.
Proteins involved¶
The full list of “agents” in the molecule_definitions.ka file is:
1%agent: ca(x)
2%agent: Glu(x)
3%agent: NMDA(x, b0, b1, state~R~RA~RA2~d1~d2~2f~2s~o, type~A~B, mgb)
4%agent: mg(x)
5%agent: AMPA(x, loc~m~b~t, ser845~u~p, ser831~u~p, b1, b2, b3, b4, state~c0~c1~c2~c3~c4~d1~d2~d3~d4~o1~o2~o3~o4)
6%agent: PMCA(x, cam)
7%agent: NCX(x)
8%agent: IMCBP(x)
9%agent: CaB(h1, h2, h3, m1)
10%agent: CaM(site1, c1, c2, n1, n2)
11%agent: CaMKII(A, B, D, cam, sub, phos, Thr286~u~p, Thr305~u~p)
12%agent: NG(cam)
13%agent: CaN(cam, sub, m1, m2)
14%agent: AC1(cam)
15%agent: AC8(cam, camLoc~none~nLobe~cLobe)
16%agent: PDE1B(x, cam, phos~u~p)
17%agent: PDE4(x, pka, phos~u~p)
18%agent: cAMP(x)
19%agent: PKA(x, A1, B1, A2, B2, type~f~r~c) #full/regulatory/catalytic
20%agent: I1(x, thr35~u~p)
21%agent: PP1(x, i1)
Rules¶
These lines from the molecule_definitions.ka demonstrate how binding cAMP to PDE4 in an unphosphorylated state:
'PDE4 binding cAMP' PDE4(x, phos~u), cAMP(x) -> PDE4(x!1, phos~u), cAMP(x!1) @ 'PDE4cAMPk+'
'PDE4 unbinding cAMP' PDE4(x!1, phos~u), cAMP(x!1) -> PDE4(x, phos~u), cAMP(x) @ 'PDE4cAMPk-'
'PDE4 destroys cAMP' PDE4(x!1, phos~u), cAMP(x!1) -> PDE4(x, phos~u) @ 'PDE4cAMPcat'
These lines demonstrate phosphorylation of PDE4 by PKAc:
'PKAc phosphorylating PDE4' PKA(x!1, type~c), PDE4(x!1, phos~u) -> PKA(x, type~c), PDE4(x, phos~p) @ 'PKAPDE4cat'
'4Ca-CaM-CaN dephosphorylating PDE4 phos' CaN(cam!2, sub!1),
CaM(site1!2, c1!_, c2!_, n1!_, n2!_), PDE4(x!1, phos~p) ->
CaN(cam!2, sub), CaM(site1!2, c1!_, c2!_, n1!_, n2!_), PDE4(x, phos~u) @ 'CaNPDE4cat'