Feature extraction¶
Overview¶
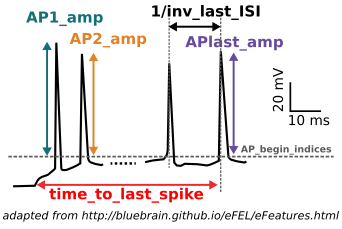
The Feature Extraction Graphical User Interface (GUI) is a web application that allows users to extract an ensemble of electrophysiological properties from voltage traces recorded upon electrical stimulation of neuronal cells (see figure on the left). The main outcome of the application is the generation of two files - features.json and protocol.json - that can be used for later model parameter optimizations.
For data selection, there are three scenarios are possible: 1) Based on her/his data permissions (see Electrophysiological data), the user selects voltage traces from the Human Brain Project (HBP) database; 2) The user selects the voltage traces from uploaded data files (through the “Upload” utility of the GUI); 3) The user selects traces from both the HBP database and her/his own uploaded file(s).
This application leverages the Python Electrophys Feature Extract Library (eFEL) and provides a friendly interface to select voltage traces (individually or collectively, based on the applied stimulus current) and features to be extracted.
Electrophysiological data¶
The Feature Extraction can only be used for voltage traces recorded during a step current stimulation protocol (see Feature Extraction). The HBP data available through the application for public use are of this type, and the user can only upload data files of the same type (see HBP database). Below is a brief description of the data formats currently in use.
HBP database¶
Publicly available data consist of electrophysiological traces provided by contributing laboratories internal and external to HBP (the contributors are shown to the user after data selection, see Trace Selection). Each data file is associated with a list of “authorized Collabs”, following the indications of the contributors. An “authorized Collab” is a collab whose team members have read access to the data. Regardless of the Collab from which they are executing the Feature Extraction web app, users who are members of, at least, one Collab authorized for a set of data, have read access to those data. Data for which the users have no authorization will not be displayed.
Contributed raw data files are in the Axon Binary Format (.abf) or the Igor Binary Format (.ibw) and each file is accompanied by a metadata.json file in which the contributors describe cell and protocol properties (e.g. cell_id, brain_region, brain_structure, stimulus_duration, …) used for data processing and visualization. The data are read through the Neo and Igorpy Python packages and converted into value arrays before use. The metadata.json files relative to the selected traces will be downloadable in a future release of the web app.
If you would like to contribute with your own data to the Feature Extraction GUI application, please send an email to bsp-support AT humanbrainproject.eu to receive further detailed information on the data format and the metadata.json template to fill in before data submission.
User’s uploaded data¶
Users can also upload and visualize their own data, which can be selected, for processing, independently or together with traces from the HBP dataset.
At this moment only the (.abf) format (Version 2 and above), produced with Molecular Devices instruments is supported for upload. To make sure that the recorded traces represent the outcome of a step current stimulation experiment, the header of the file is checked for compatibility as well. Uploading is denied when a file does not fulfill requirements.
Below is an example of a permitted protocol header, extracted from a representative .abf file with the neo package. In this example, the stimulus consisted of three periods of duration 500, 10000 and 500 ms respectively, with an initial amplitude of zero. During the central period, though, the stimulus amplitude is increased by 10.0 at every recording sweep, as indicated in the lEpochLevelInc parameter (current unit is nA).
{"dictEpochInfoPerDAC": {"0": {"0": { "fEpochInitLevel": 0.0,
"fEpochLevelInc": 0.0,
"lEpochDurationInc": 0,
"lEpochInitDuration": 500,
"lEpochPulsePeriod": 0,
"lEpochPulseWidth": 0,
"nDACNum": 0,
"nEpochNum": 0,
"nEpochType": 1,
},
"1": { "fEpochInitLevel": 0.0,
"fEpochLevelInc": 10.0,
"lEpochDurationInc": 0,
"lEpochInitDuration": 10000,
"lEpochPulsePeriod": 0,
"lEpochPulseWidth": 0,
"nDACNum": 0,
"nEpochNum": 1,
"nEpochType": 1,
},
"2": { "fEpochInitLevel": 0.0,
"fEpochLevelInc": 0.0,
"lEpochDurationInc": 0,
"lEpochInitDuration": 500,
"lEpochPulsePeriod": 0,
"lEpochPulseWidth": 0,
"nDACNum": 0,
"nEpochNum": 2,
"nEpochType": 1,
}
}
}
}
Feature Extraction¶
The Feature Extraction process precedes the generation of the features.json and protocol.json files, which are used for model parameter optimization performed through the BluePyOpt software tool (please refer to the BluePyOpt documentation for detailed explanation).
The features that the user can select for extraction are described under this link as well as the eFEL software package used to process the data. See also Hippocampal Neurons in this guidebook.
Features are computed for every trace of the chosen recordings, where a trace corresponds to a given stimulus current amplitude (indicated to the user when data are displayed). Once the extraction is finalized, the values of a feature obtained from traces belonging to an individual cell and corresponding to the same stimulation amplitude are averaged. The averages computed for all the cells, are averaged a second time by stimulus amplitude. This will generate two result files (i.e. features.json and protocols.json) per cell plus two supplementary files with the global averages.
While the eFEL extracts the features of interest from single traces (individually selected or grouped), it does not take into account any information on the cell properties, such as the cell_id needed to group the results for the generation of the above mentioned features.json and protocol.json. To perform this wrapping we used a custom Python code (please contact bsp-support AT humanbrainproject.eu for further information).
Graphical User Interface¶
The GUI guides the user through the feature extraction process in a friendly way. The web application homepage shows a brief tutorial on the usage of the interface. After the user has read (or skipped) the howto (s)he is requested to accept the Terms & Conditions for the use of the public data (see figure below).
The following is a description of the Feature Extraction process.
Trace Selection¶
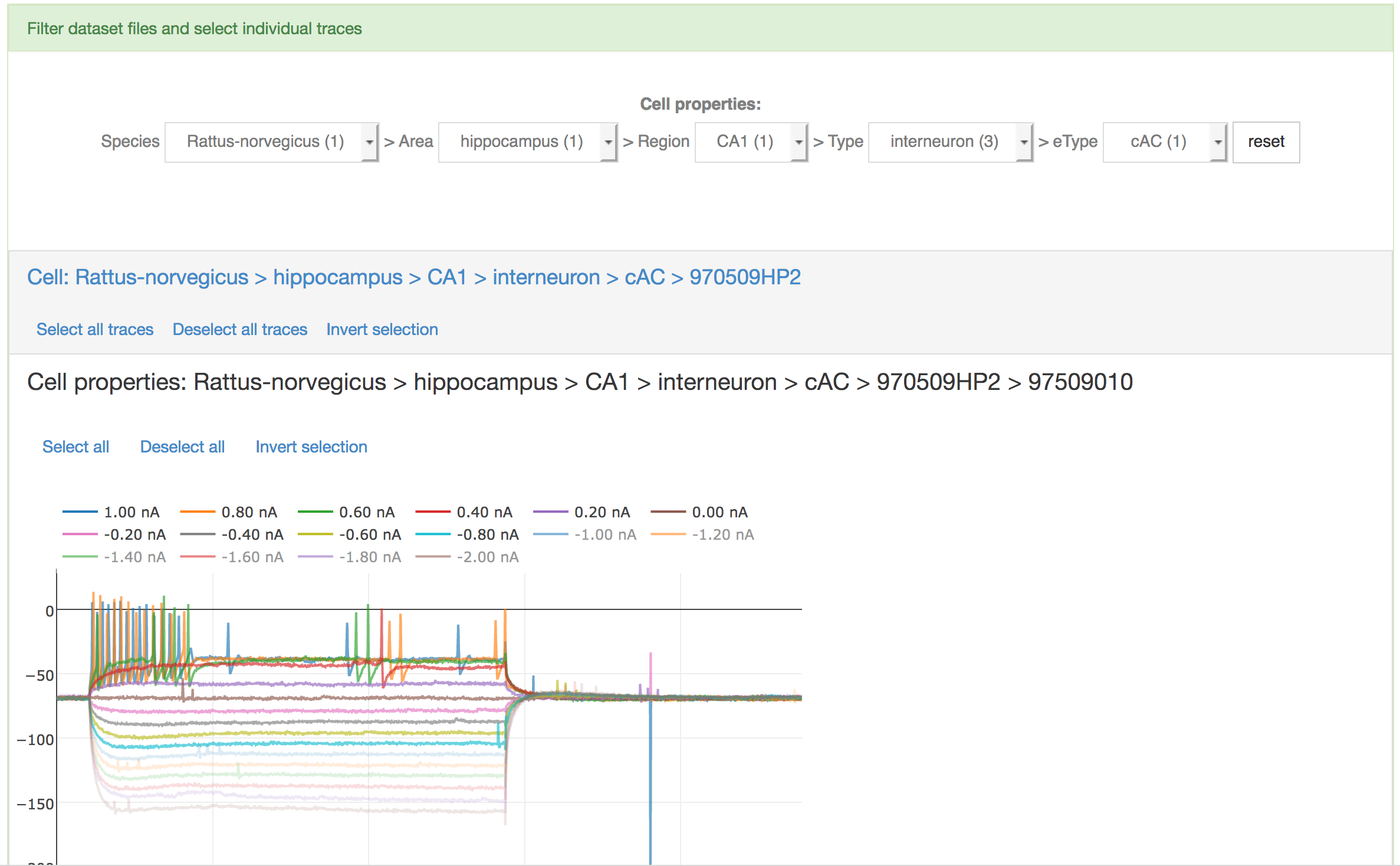
Once the trace selection page is displayed, data can be filtered from the HBP dataset by the user, by choosing the following properties of the cell from five menu lists: 1) Species; 2) Brain structure; 3) Region; 4) Type; 5) Electrical type (eType). If any of the fields is missing (e.g. the eType of the cell is not known), the “unknown” label is displayed.
When the data are loaded, the traces contained in each file are shown and data files are grouped by cell id (see figure below). The user can select individual traces by clicking on the corresponding amplitude or (s)he can select/deselect all traces in a single file. Additionally, all files (and then all traces) referring to a single cell can be selected.
Alternatively, or concurrently, users can upload their own data (see User’s uploaded data) by browsing their local storage and using the upload button (see figure below). The traces will be displayed for selection, together with the ones selected from the HBP dataset (if any).
Feature selection¶
Once the trace selection approved, the feature selection page is displayed. Features are grouped by type -spike event features, spike shape features, voltage features- and can be selected individually or collectively through the select/deselect buttons. Given the high number of features, the three types are grouped in toggle boxes (see figure below). Upon selection approval, the feature extraction process takes place as described in Feature Extraction).
Results¶
Finally, a success message is displayed and results are made available to download. The application output consists of a features.json and protocols.json files which are generated for both individual cells and the entire ensemble. These files contain the feature value averages (computed as outlined in the above section “Feature Extraction”) and the protocols adopted for the experimental recordings. These files are intended to be used for the data-driven optimization step of the Hodgkin-Huxley Neuron Builder workflow, made available to the user through the Brain Simulation Platform at the Highly Integrated Workflows page.